Shedding some light on lanternfish
 Monday, January 24, 2011 at 7:42AM
Monday, January 24, 2011 at 7:42AM  Diaphus parri, a lanternfish from the Coral Sea. Img: Adrian Flynn
Diaphus parri, a lanternfish from the Coral Sea. Img: Adrian Flynn
Adrian Flynn’s PhD is about myctophids, or lanternfishes. These are not well known to most people, but they’re probably among the most abundant animals on the planet, because they live in the largest habitat there is: the deep mid-waters of all the oceans of the world, neither near the surface nor on the bottom, what scientists call the mesopelagic zone (meso = middle, pelagic = in the water column). Lanternfish, as the name implies, produce their own light from organs arrayed on the skin of the head and body. These organs, which generate light through the action of the enzyme luciferase, allow the fish to signal each other, to find food and to disguise their own outline against the gloom from above, when seen from below. Its a neat trick, but not nearly unique in the deep pelagic zone. Indeed, it seems that just about everything down there makes light in some form or another (if you want some google fodder for that, check out Edith Widder’s work, she’s got a great talk on Ted.com too). Lanternfishes can be hard to study because they’re hard to collect in tact; they’re sort of flabby and the skin comes off very easily. But not to be deterred, Adrian is undertaking an ambitious study of how different species of lanternfishes are distributed from the tropic of Capricorn to the waters of the Antarctic - their biogeography - and also how the numbers of any given species are affected by oceanographic factors like major currents and places where deep nutrient-rich water comes up to the surface (upwelling).
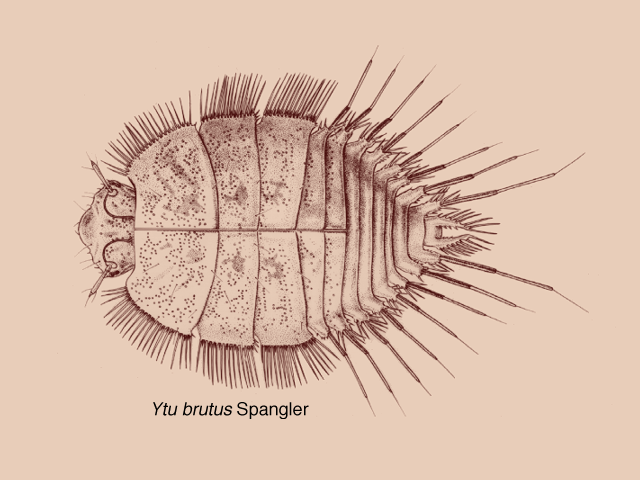
 A nice ventral view of D. parri, showing the light organs arrayed on the skin
A nice ventral view of D. parri, showing the light organs arrayed on the skin
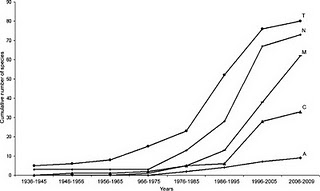
So far his results are showing that the deep pelagic zone is not as homogenous as previously thought. It seems that lanternfish distribute themselves into biogeographic zones somewhat according to latitude, but more so according the oceanographic features like major currents and landforms like islands. He’s had a couple of real eye opening results too. In one case, they observed - for the first time in Australia - a lanternfish spawning aggregation, off the coast of Cairns in far north Queensland. That’s cool, but what was a real trip was that the laternfish in question (the Dana lanternfish) was a species known only from Tasmania, almost 2,000km away! How did they get up there? How will their young get back again? On another expedition, they found lanternfish close to the surface at Macquarie Island, a remote rock in the Great Southern Ocean. The island juts up into the prevailing currents, causing upwelling that brings nutrients and lanternfishes alike well within the foraging range of penguins and seals/sea lions that nest on the island. It looks like lanternfish in this unique location are an important part of the diet of at least three penguin species as well as the pinnipeds (seals), and that’s a pretty novel discovery.
 Post a Comment |
Post a Comment |  Email Article | tagged
Email Article | tagged  Australia,
Australia,  Deep sea,
Deep sea,  biogeography,
biogeography,  lanternfish,
lanternfish,  pelagic,
pelagic,  taxonomy
taxonomy